|
BioInt
1.02.00
BioInt: An integrative biological object-oriented application framework and interpreter
|
|
BioInt
1.02.00
BioInt: An integrative biological object-oriented application framework and interpreter
|
#include <BioMultipleSequenceAlignment.h>
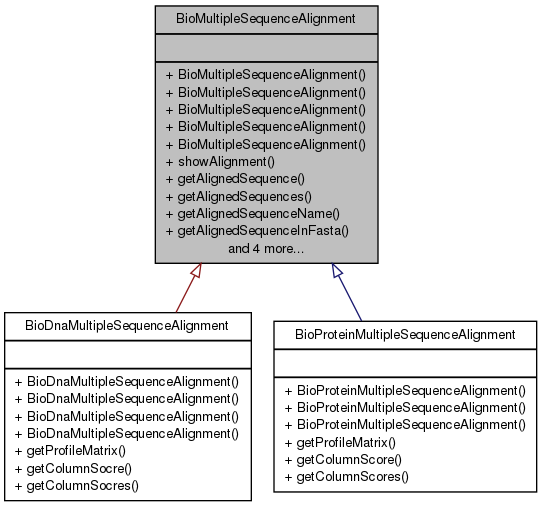
Public Member Functions | |
| BioMultipleSequenceAlignment () | |
| BioMultipleSequenceAlignment (BioMultipleFasta &) | |
| BioMultipleSequenceAlignment (BioMultipleGenBank &) | |
| BioMultipleSequenceAlignment (BioMultipleEmbl &) | |
| BioMultipleSequenceAlignment (BioMultipleSwissProt &) | |
| void | showAlignment (ostream &=cout) |
| string | getAlignedSequence (int i) |
| vector< string > | getAlignedSequences () |
| string | getAlignedSequenceName (int i) |
| BioFasta | getAlignedSequenceInFasta (int i) |
| BioMultipleFasta | getAlignedSequencesInMultipleFasta () |
| vector< char > | getColumn (int i) |
| BioMatrix | getProfileMatrix (string s) |
| int | getColumnScore (int col_index, string s) |
| vector< int > | getColumnScores (string type) |
| string BioMultipleSequenceAlignment::getAlignedSequence | ( | int | i | ) |
| string BioMultipleSequenceAlignment::getAlignedSequenceName | ( | int | i | ) |
| vector<string> BioMultipleSequenceAlignment::getAlignedSequences | ( | ) |
| vector<char> BioMultipleSequenceAlignment::getColumn | ( | int | i | ) |
| int BioMultipleSequenceAlignment::getColumnScore | ( | int | col_index, |
| string | s | ||
| ) |
| vector<int> BioMultipleSequenceAlignment::getColumnScores | ( | string | type | ) |
| void BioMultipleSequenceAlignment::showAlignment | ( | ostream & | = cout | ) |